On the nature of variation:
random, biased and directional
International conference. 3 & 4 October 2017
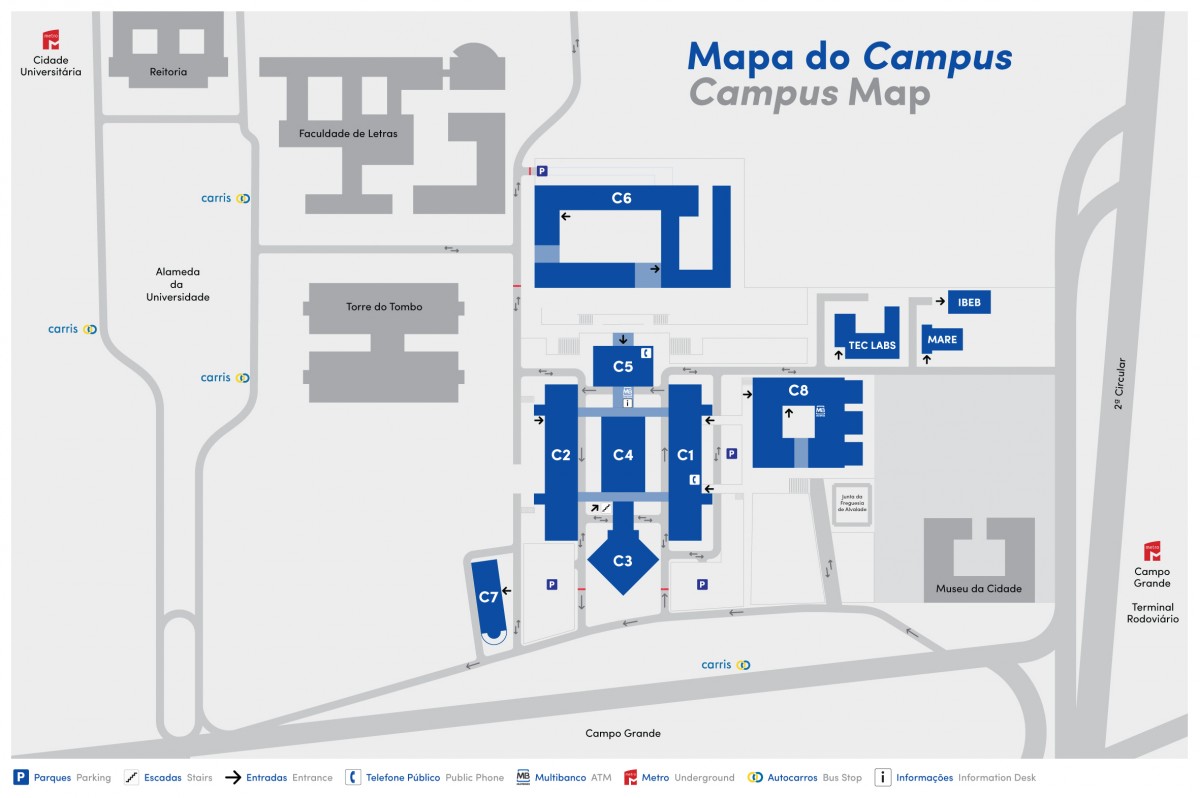
Anfiteatro da Fundação FCUL, C1 building, Faculty of Sciences
Campo Grande Campus, University of Lisbon
Adaptationism, i.e. the claim that natural selection provides a sufficient explanation for the evolution of most traits, pervades all aspects of biological thinking. The underlying assumption supporting adaptationism is that variation is somehow random, namely, that it is neither biased nor directional. This conference aims to provide an interdisciplinary context for uncovering and critically evaluating the rationale behind the hypothesis of variation randomness in the light of new developments in the evolutionary sciences (e.g., from the impact of instructive mutations in prokaryotes – CRISPR-Cas -, to mutation-biased divergence in molecular sequences, to the likely role of developmental biases in phenotypic divergence). Why was variation characterised as random in the first place? And what would be the case if either mutational or developmental biases were to impinge on the evolutionary process?
INVITED SPEAKERS: Eva Boon (Technische Universiteit Eindhoven, The Netherlands), Pietro Corsi (University of Oxford, UK and EHESS, Paris, France), Leonore Fleming (Utica College, USA), Philippe Huneman (IHPST; CNRS/ Université Paris 1 Sorbonne), Gerd Müller (Universität Wien, Austria), Sahotra Sarkar (University of Texas at Austin, USA and Presidency University, Kolkata, India), Arlin Stoltzfus (Institute for Bioscience and Biotechnology Research, NIST, USA).
CALL FOR ABSTRACTS AND PROGRAMME
Call for Abstracts
The present call for abstracts is directed to philosophers of biology, philosophers of science, historians of biology, evolutionary and theoretical biologists. The aim of the conference is to provide an interdisciplinary context for evaluating the historical, empirical and theoretical dimensions of the debate concerning the nature of variation and the proper role for variation in evolutionary explanations, how the debate has unfolded and how it shapes current evolutionary thinking in the life sciences.
- How useful is the doctrine of variational randomness? And how should it be characterised?
- In what sense would the existence of processes of generation of biased or directional variation challenge (empirical) adaptationism?
- Is phenotypic variation random in the same sense as genomic variation?
- Are all processes of genomic change random in the same sense?
- How can evolutionary thinking benefit from concepts of randomness used in other sciences (e.g., mathematics, physics)?
- To what extent do indeterministic processes play a role in mutation?
- What mutation systems other than CRISPR-Cas violate current notions of randomness?
- How can evolutionary thinking benefit from an analogy with cultural processes of variation-generation?
Programme
OCTOBER 3
9.45-10.00 | Welcome
10.00-11.00 | Pietro Corsi
University of Oxford, UK and EHESS Paris, France
The inheritance of acquired characteristics:
Darwin vs Lamarck, or Darwin and Lamarck?
11.00-11.30 | David Suárez Pascal
Universidad Nacional Autónoma de México, México
A function-based analysis of randomness in evolutionary change
11.30-12.00 | Coffee break
12.00-13.00 | Leonore Fleming
Utica College, USA
Nothing in biology makes sense except in the light of variation
13.00-13.30 | Pablo Razeto-Barry & Davide Vecchi
Instituto de Filosofía y Ciencias de la Complejidad, Chile | Universidade de Lisboa, Portugal
Genomic change randomness: a net-fitness-effect account
13.30-15.00 | Lunch
15.00-16.00 | Eva Boon
Technische Universiteit Eindhoven, The Netherlands
In which sense is lateral gene transfer random, and why does it matter?
16.00-16.30 | Coffee break
16.30-17.00 | Lorenzo Baravalle
Universidade Federal do ABC, Santo André, São Paulo, Brazil
The cultural selection of chance
17.00-18.00 | Philippe Huneman
IHPST, CNRS, Université Paris 1 Sorbonne, France
Distinguishing types of variation and assessing challenges to the Modern Synthesis
18.00-18.30 | Discussion
OCTOBER 4
10.00-11.00 | Gerd Müller
Universität Wien, Austria
The morphogenetic basis of discontinuous variation
11.00-11.30 | James DiFrisco
Konrad Lorenz Institute for Evolution and Cognition Research, Austria
Variation across levels of organization:
insights from evo-devo
11.30-12.00 | Coffee break
12.00-12.30 | Laura Nuño de la Rosa & Cristina Villegas
University of the Basque Country, Spain | Universidad Complutense de Madrid, Spain
Possible forms and possible adaptations, or how evolvability challenges random variation
12.30-13.30 | Arlin Stoltzfus
Institute for Bioscience and Biotechnology Research, NIST, USA
A new kind of variational cause, and some implications
13.30-15.00 | Lunch
15.00-15.30 | Julian Zhunping Xue
McGill University, Canada
From mutation rates to eukaryotic organization:
how variation drives evolution even in regimes of strong selection
15.30-16.00 | Predrag Šustar & Zdenka Brzović
University of Rijeka, Croatia
Gene duplication between mechanism and random process
16.00-16.30 | Coffee break
16.30-17.00 | Giorgio Airoldi
Universidad Nacional de Educación a Distancia, Madrid, Spain
Evolution without selection: how shrinking the scope of fitness as the explanatory driver of evolutionary changes can reconcile adaptationism and pluralism
17.00-18.00 | Sahotra Sarkar
University of Texas at Austin, USA and Presidency University, Kolkata, India
Blind variation and evolutionary explanation
18.00-18.30 | Discussion
Registration free but mandatory: sdmarco@fc.ul.pt
Download programme
Download Book of Abstracts
INVITED SPEAKERS
(Technische Universiteit Eindhoven, The Netherlands)
https://www.tue.nl/en/university/departments/industrial-engineering-innovation-sciences/the-department/staff/detail/ep/e/d/ep-uid/20157707/

(University of Oxford, UK and EHESS, Paris, France)
http://www.history.ox.ac.uk/people/professor-pietro-corsi

Leonore Fleming
(Utica College, USA)
http://www.utica.edu/academic/hhs/philosophy/philosophyfaculty.cfm?featureaction=details&id=E5E74582-F2F0-C47C-307B692FB0D5CF5C

(Institut d’Histoire et de Philosophie des Sciences et des Techniques;
CNRS/ Université Paris 1 Sorbonne)
http://philippehuneman.wordpress.com


(University of Texas at Austin, USA and Presidency University, Kolkata, India)
https://liberalarts.utexas.edu/philosophy/faculty/sarkars1

ORGANISATION
Convenors:
Elena Casetta, Silvia Di Marco, Jorge Marques da Silva, Carina Vieira da Silva, Davide Vecchi
Funded by the FCT (Fundação para a Ciência e a Tecnologia) through the Philosophy of the Life Sciences Group of the Centro de Filosofia das Ciências da Universidade de Lisboa (strategic project UID/FIL/00678/2013) and the R&D Project Biodecon (PTDC/IVC-HFC/1817/2014).
Description and rationale of the conference
Adaptationism pervades all aspects of biological thinking. Very generally, adaptationism could be characterised as the claim that natural selection provides a sufficient explanation for the evolution of most traits. This view (labelled empirical adaptationism) enshrines natural selection as the principle of the order of nature, with the consequence that other evolutionary processes are thought as having a negligible influence on trait evolution (Orzack & Forber 2010, https://plato.stanford.edu/entries/adaptationism/#EmpAda). Needless to say, such strong empirical hypothesis has not remained unchallenged. One crucial challenge to adaptationism stems from recent genomic analyses (Sarkar 2014) suggesting that non-adaptive processes such as drift and mutation dominate genome evolution:
Numerous aspects of genomic architecture, gene structure, and developmental pathways are difficult to explain without invoking the nonadaptive forces of genetic drift and mutation. (Lynch 2007a, p. 8597).
Studies of molecular evolution have reshaped the debate concerning the nature of evolution in many respects. On the one hand, credible neutralist hypotheses accounting for both genomic and phenotypic evolution have been proposed (Lynch 2007b, Gray et al. 2010, Stoltzfus 2012). On the other, the nature of the processes of genomic change has been uncovered in its variety and complexity: a plethora of different processes of genomic change with different mechanistic features exists, varying from small sequence changes to gene duplications and entire genome doublings (for a review see Koonin 2009). One corollary of these developments is that, if genomic change is not random, it might bias evolution.
A different challenge to adaptationism stems from studies of phenotypic evolution suggesting that developmental processes co-determine the direction of evolution:
Developmental processes, operating through developmental bias and niche construction, share with natural selection some responsibility for the direction and rate of evolution …. (Laland et al. 2015, p. 2)
Particularly interesting is the suggestion that phenotypic variation is not random, an idea clarified with characteristic acumen by West-Eberhard (2005, p. 6543):
Insofar as phenotypic novelties arise from adaptive developmental plasticity, they are not ‘random’ variants, because their initial form reflects adaptive responses with an evolutionary history, even though they are initiated by mutations or novel environmental factors that are random with respect to (future) adaptation.
Developmental bias, for instance, makes the appearance and fixation of certain phenotypic variants more probable than others (Lange & Müller 2015, Lange et al. 2014, Müller 2007, Peterson & Müller 2016). A corollary of this theoretical perspective is that, if phenotypic change is not random, it might bias evolution.
One way to capture the common theoretical thread of these two challenges is the following: genomic and phenotypic variation bias evolutionary outcomes because they are not “random”. This conference aims to provide an interdisciplinary context for uncovering and critically evaluating the rationale behind the hypothesis of variation randomness in the light of new developments in the evolutionary sciences. Why was variation characterised as random in the first place? And what would be the case if either mutational or developmental biases were to impinge on the evolutionary process?
Why is variation considered “random”?
Textbooks in evolutionary biology help to understand the rationale behind the idea of randomness. For instance, Ridley (2004, pp. 88-9) stresses the idea of uncoupling of variation and selection:
A basic property of Darwinism is that the direction of evolution, particularly of adaptive evolution, is uncoupled from the direction of variation. When a new recombinant or mutant genotype arises, there is no tendency for it to arise in the direction of improved adaptation. Natural selection imposes direction on evolution, using undirected variation.
Futuyma (2005, p. 179) makes the same point by invoking the spectre of Lamarckism:
The argument that adaptively directed mutation does not occur is one of the fundamental tenets of modern evolutionary theory. If it did occur, it would introduce a Lamarckian element into evolution, for organisms would then acquire adaptive hereditary characteristics in response to their environment. Such neo-Lamarckian ideas were expunged in the 1940s and 1950s by experiments with bacteria in which spontaneous, random mutation followed by natural selection, rather than mutation directed by the environment, explained adaptation.
In brief, if the processes generating genomic and phenotypic variation were not random, the limitations of the adaptationist explanation of biological adaptedness, diversity and complexity would be revealed.
Why has mutation been characterised as a “random” process?
As Futuyma argues in the previous quote, the hypothesis of the randomness of mutation dominating evolutionary biology probably took form following the famous fluctuation test by Luria and Delbrück in 1943. Luria (1984) claimed that he devised the experiment by watching a colleague playing on a slot machine, realizing that “… slot machines and bacterial mutations have something to teach each other.” Luria and Delbrück thought that if bacterial mutations were induced by phage sensitivity, they would be evenly distributed among the cultures. But if phage resistance were to emerge spontaneously (i.e., if resistance is not environmentally induced), the outcome would be a pattern of clustered resistant colonies, analogous to slot machine jackpots. The latter distribution was observed, concluding that mutation is a spontaneous process. Interestingly, in Luria’s (1984, p. 80) autobiography, he switched between an epistemological (i.e., mutation as an unpredictable slot-machine’s jackpot) and an ontological (i.e., mutation as a process analogous to radioactive decay) interpretation of randomness. The epistemological interpretation is compatible with assuming that mutagenesis is a deterministic process whose triggering causes are unknown. In contrast, if mutation is analogized to radioactive decay, then it might be regarded as indeterministic in nature. It is certainly possible that some mutagenic processes are influenced by quantum indeterminacy (Stamos 2001), even though it would be preposterous to conclude that all processes of variation generation are random in this very strong sense. Nonetheless, textbooks in evolutionary biology borrow from this latter narrative. For instance, Futuyma (2005, p. 178) claims that:
The spontaneous process of mutation is stochastic rather than deterministic.
The biological consequences of endorsing nonchalantly such a view are clear enough: if mutagenesis is indeterministic in character, why should significant generalisations be expected concerning its role in evolution? Evolutionary biology should focus on studying selection instead. Adaptationism would be reinforced.
The evolutionary sense of randomness
Understanding the reasons why the obviously inappropriate term “random” was chosen to characterise mutation is an interesting topic for historical research (for a history of mutation see Carlson 2011). The term “random” could for instance refer to spontaneous, indeterministic, stochastic or unbiased changes. All these senses of the term are different. It is however important to emphasise that in the evolutionary literature the thesis of the randomness of variation is qualified and used in a more specific – though still difficult to characterise precisely – sense. Variation is random only with respect to adaptation. Thus, the basic idea is only that variation is not directional (i.e., adaptive). For instance, Futuyma (2005, pp. 178-9) claims that:
… mutation is random in the sense that the chance that a particular mutation will occur is not influenced by whether or not the organism is in an environment in which the mutation would be advantageous. That is, the environment does not induce adaptive mutations. Indeed, it is hard to imagine a mechanism whereby most environmental factors could direct the mutation process by dictating that just the right base pair changes should occur.
Analogously, Ridley (2004, pp. 88-9) asks to consider the scenario in which a virus encounters a new environment and claims that:
If we reflect on the kind of mechanism that would be needed, it becomes clear that an adaptively directed mutation would be practically impossible. The virus would have to recognize that the environment had changed, work out what change was needed to adapt to the new conditions, and then cause the correct base changes.
This popular view of mutation as random (with respect to adaptation and phenotypic improvement) – complemented with the view that selection is the sole cause of creativity and direction in evolution – was generalised to encompass indiscriminately all processes of variation generation. Furthermore, it was also transferred to other fields of research. For instance, Popper (1972) proposed that even scientific evolution is governed by partially chance-like conjectures (analogous to mutation) and refutations (analogous to selection), while Campbell (1974) proposed that blind variation followed by selective retention is the fundamental process of both biological and cultural evolution (see also Stein & Lipton 1989, Mesoudi 2008, Kronfeldner 2010, Vecchi & Baravalle 2015).
Is the evidence for mutation randomness sound?
The foundational experimental work in microbiology of the 1940s and 1950s was not sufficient to dismiss the possibility of directional mutagenesis. Many experiments – such as Luria’s – subjected the organisms to lethal factors from which they could not recover and exhibit adaptive responses. More generally, the hypothesis that mutation is random solidified despite a profound ignorance of the molecular and biochemical nature of the cellular machinery (e.g., DNA repair). Thus, it could be argued that the view that mutations (indeed all processes of genomic change) are random with respect to adaptation has crystallised without a sound evidential basis. As Shapiro (2011, p. 79) puts it, there exists:
… an unfortunate tendency in evolutionary studies … to make sweeping assertions about what can and cannot happen based on relatively limited experimental observations.
In this sense, it is significant that, in the last quotations, both Futuyma and Ridley make reference to theoretical reasons for dismissing the possibility that mechanisms of directional mutagenesis might exist. It is also interesting to note that it was eventually a new wave of experimental work that reinvigorated the debate concerning the randomness of mutation. New empirical evidence uncovered from the 1980s suggests that many unicellular organisms might be able to respond adaptively to environmental conditions (e.g., Shapiro 1984; Cairns et al. 1988, Hall 1990, 1998, Wright 2000, Rosenberg 2001). This kind of research led to an important theoretical discussion as well as to many alternative interpretations of the empirical evidence (see for instance Keller 1992, Sober 1992, Lenski and Mittler 1993, Jablonka and Lamb 1995, Cairns 1998, Brisson 2003, Sarkar 2005).
Where do we stand now?
Recent experimental research in molecular biology has transformed the debate about the nature of mutagenesis by unravelling many mechanistic details of a variety of mutagenic processes. What can be safely claimed today is that mutagenesis is sometimes environmentally induced, that it is biased in a number of ways and that it is sometimes a regulated process. What remains debatable is whether there exist mechanisms that regulate mutagenesis so as to increase the probability of beneficial mutations vis a vis deleterious ones. As it can be expected in the case of theoretically complex disputes, opinions diverge. The debate has been recently enriched by a series of contributions both by philosophers of biology (Merlin 2010, Sober 2014, Baravalle & Vecchi 2016) and biologists (Jablonka & Lamb 2006, Koonin and Wolf 2009, Stoltzfus & Yampolsky 2009, Shapiro 2011, Razeto-Barry & Vecchi 2017). But the debate concerning randomness is not solely confined to mutagenesis as it encompasses all processes of genomic and phenotypic change.
Why is this debate important?
The debate concerning the nature of variation is of fundamental importance because it allows evaluating adaptationist claims in two senses. If variation is not random, then it is either biased or directional. If the processes of generation of genomic and phenotypic variation were biased, then evolutionary outcomes would depend not solely on natural selection, but on the nature of these biases: mutational and developmental biases would therefore be processes of fundamental evolutionary importance. If the processes of generation of genomic and phenotypic variation were directional, then evolutionary outcomes would depend not solely on natural selection, but on the capacities of the organisms to respond adaptively to environmental challenges. Koonin and Wolf (2009) make this point in reference to the nature of genomic variation:
….the radical reappraisal of the nature of genomic variation and the realization that much of this variation is adaptive, thus apparently eliminating the conflict between the Lamarckian and Darwinian scenarios, is a veritable, although underappreciated paradigm shift in modern biology.
The extent to which the same argument applies to phenotypic variation, and the extent to which biased and directional variation direct evolution, remains open to debate. Indeed, the ultimate aim of this conference is to explore these issues.
Bibliography
Baravalle L & Vecchi D. (2016). Beyond blindness: On the role of organism and environment in trial generation. Studies in History and Philosophy of Science Part C: Studies in History and Philosophy of Biological and Biomedical Sciences 60:25-34
Brisson D. (2003). The directed mutation Controversy in an Evolutionary Context. Critical Reviews in Microbiology, 29, 25–35
Cairns J. (1998) Mutation and cancer: the antecedents to our studies of adaptive mutation. Genetics 148:1433–1440
Cairns J, Overbaugh J, Miller S. (1988) The origin of mutants. Nature 335:142
Carlson EA. (2011). Mutation: The History of an Idea from Darwin to Genomics. Codld Spring Harbour Laboratory Press
Campbell DT. (1974). Unjustified variation and selective retention in scientific discovery. In F.J. Ayala & T. Dobzhansky (Eds.), Studies in the philosophy of biology (pp. 139–162). New York: Macmillan.
Futuyma DJ. (2005). Evolution. Sinauer Associates.
Gray MW, Lukes J, Archibald JM, Keeling PJ, Doolittle WF. (2010) Cell biology. Irremediable complexity? Science 330(6006):920–921.
Hall BJ. (1990). Spontaneous point mutations that occur more often when advantageous than when neutral. Genetics 126, 5–16.
Hall BJ. (1998). Adaptive mutagenesis: a process that generates almost exclusivelybeneficial mutations. Genetica 102/103, 109–125.
Jablonka E, Lamb M (1995) Epigenetic inheritance and evolution: the Lamarckian dimension. Oxford University Press, Oxford
Jablonka E & Lamb M. (2006). Evolution in four dimensions. Cambridge, MA: MIT Press
Keller EF (1992) Between language and science: the question of directed mutation in molecular genetics. Perspect Biol Med 35:292–306
Koonin EV. (2009). Darwinian evolution in the light of genomics. Nucleic Acids Research, 37(4):1011–1034.
Koonin EV & Wolf YI. (2009). Is evolution Darwinian or/and Lamarckian? Biology Direct, 4(42). (http://www.biologydirect.com/content/4/1/42)
Kronfeldner M. (2010). Darwinian “blind” hypothesis formation revisited. Synthese, 175, 193e218.
Laland K, Uller T, Feldman MW, Sterelny K, Müller GB, Moczek M, Jablonka E, Odling-Smee J. (2015). The extended evolutionary synthesis: Its structure, assumptions and predictions. Proceedings of the Royal Society B (http://rspb.royalsocietypublishing.org/content/282/1813/20151019).
Lange A, Müller GB. 2014 Polydactyly in development, inheritance, and evolution. The Quarterly Review of Biology 92(1): 1-38.
Lange A, Nemeschkal HL, Müller GB. 2014. Biased polyphenism in polydactylous cats carrying a single point mutation: The Hemingway model for digit novelty. Evol Biol 41:262–275.
Lenski R & Mittler JE. (1993). The directed mutation controversy and Neo-Darwinism. Science, 259, 188e194
Luria SE (1984) A slot machine, a broken test tube. Harper and Row, New York
Luria SE & Delbrück M. (1943). Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28(6), 491–511
Lynch M. (2007a). The frailty of adaptive hypotheses for the origins of organismal complexity. Proc Natl Acad Sci USA 104(Suppl 1):8597–8604.
Lynch M. (2007b). The Origins of Genome Architecture (Sinauer, Sunderland, MA).
Merlin F. (2010). Evolutionary chance mutation: a defense of the modern synthesis’ consensus view. Philosophy and Theory in Biology 2, e103
Mesoudi A. (2008). Foresight in cultural evolution. Biology and Philosophy, 23(2), 243–255.
Müller GB. 2007. Evo-devo: Extending the evolutionary synthesis. Nat Rev Genet 8:943–949.
Orzack SH, Forber P (2010). Adaptationism. The Stanford Encyclopedia of Philosophy. (https://plato.stanford.edu/entries/adaptationism/#EmpAda)
Peterson, T. & Müller, GB. (2016). Phenotypic Novelty in EvoDevo: The Distinction Between Continuous and Discontinuous Variation and Its Importance in Evolutionary Theory. Evolutionary Biology 43:314–335
Popper KR. (1972). Objective knowledge. Oxford: Oxford University Press.
Razeto-Barry P and Vecchi D. (2017). Mutational randomness as conditional independence and the experimental vindication of mutational Lamarckism. Biol Rev Camb Philos Soc, (92-2):673–683
Ridley M. (2004) Evolution (3rd Edition). Blackwell Science.
Rosenberg SM. (2001). Evolving responsively: adaptive mutation. Nature Reviews Genetics 2(7), 504–515.
Sarkar S. (2005). Molecular Models of Life. MIT Press, Cambridge
Sarkar S. The genomic challenge to adaptationism. British Journal for the Philosophy of Science. 2014
Shapiro JA. (1984) Observations on the formation of clones containing araB-lacZ cistron fusions. Molec Gen Genet 194:79–90
Shapiro JA. (2011). Evolution: A view from the 21st century. Upper Saddle River, NJ: FT Press Science.
Sober E. (1992). Models of cultural evolution. In P. Griffiths (Ed.), Trees of Life: Essays in philosophy of biology (pp. 17-39). Dordrecht: Kluwer
Sober E. (2014). Evolutionary Theory, Causal Completeness, and Theism: the Case of ’Guided’ Mutation, in Thompson RP & Walsh D eds Evolutionary Biology: Conceptual, Ethical, and Religious Issues pages 31–44. Cambridge University Press, Cambridge, 2014
Stamos DN. (2001). Quantum Indeterminism and Evolutionary Biology. Philosophy of Science, 68(2):164, 2001.
Stein E & Lipton P. (1989). Where guesses come from: Evolutionary epistemology and the anomaly of guided variation. Biology and Philosophy, 4(1), 33–56.
Stoltzfus A. (2012). Constructive neutral evolution: exploring evolutionary theory’s curious disconnect. Biol Direct, 7(1):35.
Stoltzfus, A. & Yampolsky, LY. (2009). Climbing Mount Probable: Mutation as a Cause of Nonrandomness in Evolution. Journal of Heredity 100(5), 637-647.
Vecchi, D. & Baravalle, L. A soul of truth in things erroneous: Popper’s “amateurish” evolutionary philosophy in light of contemporary biology. History and Philosophy of the Life Sciences, 36:525-545, 2015.
West-Eberhard, M. J. (2005). Developmental plasticity and the origin of species differences. PNAS, 102(Suppl. 1), 6543–6549.
Acknowledgements
Artwork: Sara Fuentes
Admin honcho: Ana Vilar Bravo
Website: CFCUL / Ricardo Pestana